
Analyze uncharted marine ecosystems and uncover hidden patterns of ocean life through critical underwater sound analysis. The ocean's soundscape teems with enigmatic noises and undiscovered phenomena. For instance, the intriguing "biotwang" sound, recently identified by the U.S. National Oceanic and Atmospheric Administration (NOAA) as originating from the elusive Bryde's whales, highlights the persistent challenge of continually identifying novel song types and species classifications.
Google possesses a robust history of collaborating with researchers on bioacoustics applications for whale monitoring and conservation. This includes our foundational AI models for detecting humpback whale vocalizations and the 2024 release of our comprehensive multi-species whale identification model. To accelerate discoveries, Google's AI approach for bioacoustics now focuses on enabling faster connections between new findings and actionable scientific insights at scale. In August 2025, Google DeepMind introduced Perch 2.0, a revolutionary bioacoustics foundation model. Surprisingly, despite training exclusively on terrestrial animal vocalizations, Perch 2.0 demonstrates exceptional performance as an embedding model for transfer learning in marine acoustic analysis.

The beluga whale, an essential indicator species in the marine environment. (Credit: Lauren Harrell)
Our recent publication, “Perch 2.0 transfers 'whale' to underwater tasks,” a joint effort by Google Research and Google DeepMind presented at the NeurIPS 2025 workshop on AI for Non-Human Animal Communications, meticulously details these findings. This bioacoustics foundation model, primarily trained on avian vocalizations, effectively facilitates and scales insights into underwater marine ecosystems, specifically for classifying whale vocalizations. We also provide an accessible, end-to-end tutorial in Google Colab demonstrating our agile modeling workflow. This guide empowers users to leverage Perch 2.0 for creating custom whale vocalization classifiers using the NOAA NCEI Passive Acoustic Data Archive via Google Cloud.
Mastering Bioacoustics Classification with AI
When an established classification model, such as our multi-species whale identifier, possesses the requisite labels and performs effectively on a researcher's dataset, it can be deployed directly to generate scores and classifications for their audio data. However, to develop novel custom classifiers for newly discovered sounds or to enhance accuracy with new data, we employ transfer learning. This sophisticated technique significantly reduces the computational demands and experimental overhead compared to building a model from the ground up.
In bioacoustics transfer learning, the pre-trained model, exemplified by Perch 2.0, generates embeddings for discrete audio windows. These embeddings condense extensive audio data into compact feature arrays, serving as input for a straightforward classifier. Creating a new custom model for any labeled audio dataset involves applying the pre-trained model to extract embeddings, which then act as input features for a logistic regression classifier. Consequently, instead of learning the extensive parameters of a deep neural network, we exclusively train new parameters for the final logistic regression stage, yielding substantial efficiency gains in both researcher time and computational resources.
Rigorous Performance Evaluation
We meticulously evaluated Perch 2.0 using a few-shot linear probe methodology across diverse marine acoustic tasks, including differentiating between various baleen whale species and distinct killer whale subpopulations. Its performance was benchmarked against established pre-trained models available in our Perch Hoplite repository, designed for agile modeling and transfer learning. These models include Perch 2.0, Perch 1.0, SurfPerch, and the multi-species whale model.
For evaluating underwater data, we utilized three comprehensive datasets: NOAA PIPAN, ReefSet, and DCLDE. Each dataset offers unique insights into marine bioacoustics:
- NOAA PIPAN: This meticulously annotated subset of the NOAA NCEI Passive Acoustic Data Archive originates from NOAA Pacific Islands Fisheries Science Center recordings. It incorporates labels previously used in our whale models and introduces new annotations for baleen species, including common minke whale, humpback whale, sei whale, blue whale, fin whale, and Bryde’s whale.
- ReefSet: Developed specifically for SurfPerch model training, this dataset incorporates data annotations from the Google Arts and Culture project, "Calling in Our Corals." It features a diverse range of bioacoustic elements, including general reef noises (croaks, crackles, growls), specific species/genera classifications (e.g., damselfish, dolphins, and groupers), and anthropomorphic noise and wave signatures.
- DCLDE: This dataset undergoes evaluation using three distinct label sets:
- Species: To discern between killer whales, humpbacks, abiotic sounds, and unidentified underwater noises (acknowledging some uncertainty in killer whale and humpback labels).
- Species Known Bio: Focusing on confirmed labels for killer whales and humpbacks.
- Ecotype: To differentiate between various killer whale subpopulations (ecotypes), encompassing Transient/Biggs, Northern Residents, Southern Residents, Southeastern Alaska killer whales, and offshore killer whales.
Within this evaluation protocol, for each target dataset with available labeled data, we extract embeddings from candidate models. We then select a fixed number of examples per class (4, 8, 16, or 32) and train a simple multi-class logistic regression model using these embeddings. The resulting classifier is employed to compute the area under the receiver-operating characteristic curve (AUC_ROC), where values approaching 1 signify a superior capability in distinguishing between classes. This process effectively simulates the application of a pre-trained embedding model to create a custom classifier from a limited set of labeled examples.
Our findings conclusively demonstrate that augmenting the number of examples per class consistently enhances performance across all evaluated models. This trend holds true for all models except on the ReefSet data, where high performance is achieved even with just four examples per class for every model, with the exception of the multi-species whale model. Significantly, Perch 2.0 consistently achieves either the top or second-best performance across all datasets and sample sizes.
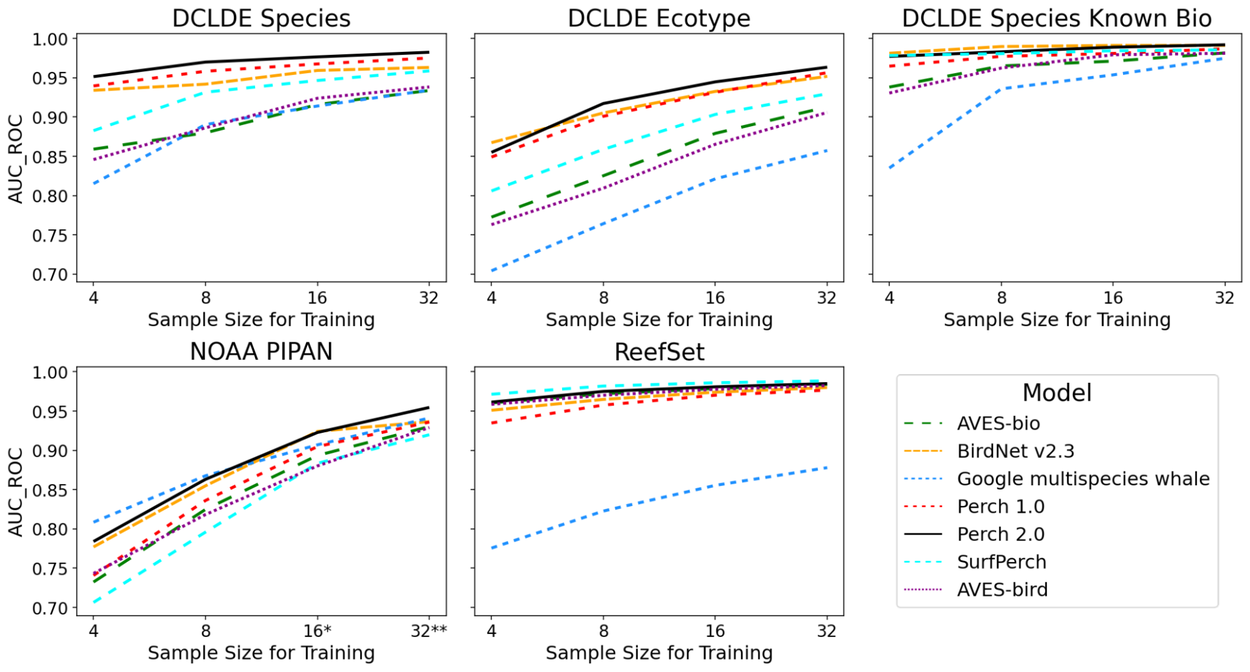
Comparative performance analysis of trained models on marine datasets, varying the number (k) of training examples per-class. Higher AUC_ROC values signify enhanced classification accuracy. *Class “Bm” was omitted for k = 16; **Classes ‘Bm’ and ‘Be’ were omitted for k = 32 in the NOAA PIPAN dataset.
Furthermore, we benchmark Perch 2.0 against embeddings derived from AVES-bird and AVES-bio (transformer bioacoustics models from the Earth Species Project trained on avian and general biological sounds, respectively), alongside BirdNet v2.3 from the K. Lisa Yang Center for Conservation Bioacoustics at the Cornell Lab of Ornithology. Perch 2.0 demonstrates superior performance over AVES-bird and AVES-bio on the majority of underwater acoustic tasks. Notably, several other pre-trained models, though not specifically trained on underwater audio, also exhibit strong performance capabilities.
Unraveling the 'Whale' in Perch 2.0's Performance
We propose several key factors contributing to Perch 2.0's remarkable transfer performance from terrestrial birds to underwater acoustics. Firstly, prior research consistently indicates that larger models trained on extensive datasets exhibit superior generalization capabilities. This allows our bioacoustics model to effectively classify sounds for species and acoustic phenomena not explicitly included in its training data. Secondly, the inherent challenge of distinguishing highly similar bird calls, often referred to as the “bittern lesson,” compels the model to learn nuanced acoustic features. These detailed features subsequently prove invaluable for diverse bioacoustics tasks. For instance, distinguishing the subtly different “coo” sounds of North America's 14 dove species requires the model to extract discriminative features that can then aid in differentiating other sound classes. Finally, the transfer of acoustic features across distinct species groups may also stem from convergent evolution in sound production mechanisms. A variety of species, including birds and marine mammals, have independently evolved similar methods for generating sound.
An effective bioacoustics model produces embeddings that are both informative and linearly separable for the target classes being analyzed. To illustrate this, we visualize a summary of embeddings from each model using t-distributed stochastic neighbor embedding (tSNE), where distinct colors denote different classes. Highly informative models reveal clearly delineated clusters for each class, whereas less informative models show greater intermingling between classes. While most models exhibit discernible clusters for Southern Resident killer whales (KW_SRKW) and Southern Alaskan Residents (KW_SAR), the embeddings for Northern Resident killer whales (KW_NRKW), Transient killer whales (KW_TKW), and Offshore killer whales (KW_OKW) often overlap in models like AVES-bio, AVES-bird, and SurfPerch. In contrast, these killer whale ecotypes are much more distinctly separated in models such as BirdNet v2.3 and Perch 2.0.
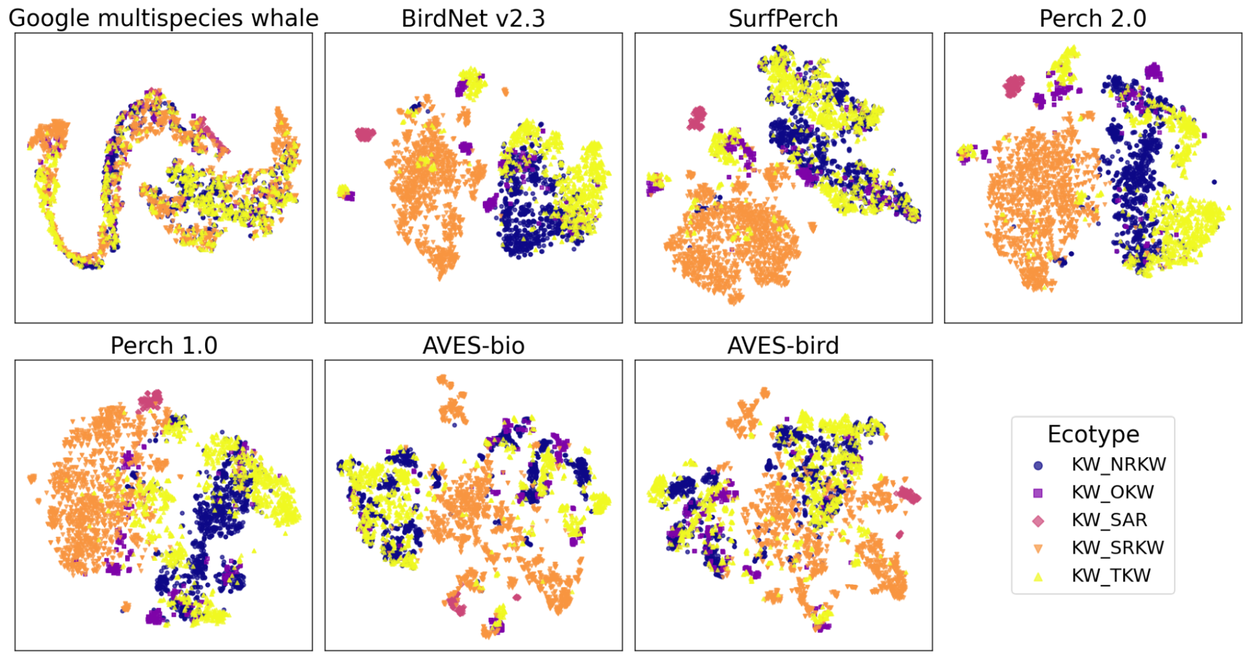

tSNE visualizations of model embeddings on the DCLDE 2026 Ecotype dataset, differentiating five killer whale (orca) ecotype variants. Plots were generated using sci-kit learn PCA and tSNE libraries, with embeddings initially projected to 32-dimensional vectors before tSNE application.
Charting the Future of Bioacoustic Discovery
The Google DeepMind Perch team, in close collaboration with Google Research and external partners, has spearheaded an agile modeling methodology for bioacoustics. This innovative approach enables the creation of custom classifiers from a minimal set of labeled examples within just a few hours. To provide comprehensive support for both Google Research collaborators and the broader cetacean acoustics community, we have developed an end-to-end demonstration. This demo facilitates working with NOAA data from the Passive Acoustic Archive dataset hosted on Google Cloud, building upon and updating our previous tutorials with the more efficient Perch Hoplite databases for managing embeddings.
Acknowledgments
The Perch Team, instrumental in developing the Perch 2.0 model and part of Google DeepMind, comprises Tom Denton, Bart van Merriënboer, Vincent Dumoulin, Jenny Hamer, Isabelle Simpson, Andrea Burns, and Lauren Harrell (Google Research). We extend our sincere gratitude to Ann Allen (NOAA Pacific Islands Fisheries Center) and Megan Wood (Saltwater Inc., supporting NOAA) for providing crucial additional annotations for the NOAA PIPAN dataset. Special thanks are also due to Dan Morris (Google Research) and Matt Harvey (Google DeepMind) for their invaluable contributions.


