
Introducing TRINDs: a comprehensive dataset and robust benchmarking pipeline engineered with synthetic personas to rigorously assess and elevate Large Language Model (LLM) capabilities specifically for tropical and infectious disease domains.
Large language models (LLMs) demonstrate significant promise in advancing medical and health question-answering, excelling across diverse health-related assessments and data formats. Google Research is at the vanguard of enhancing LLM utility in healthcare, evidenced by our pioneering work on Med-Gemini, MedPaLM, AMIE, Multimodal Medical AI, and the release of advanced evaluation tools. In underserved regions, LLMs can revolutionize clinical decision support, improve diagnostic precision, broaden access, facilitate multilingual clinical guidance, and enhance community health education. However, current medical benchmarks may not fully capture LLM generalization capabilities when faced with distribution shifts in disease types, region-specific medical nuances, and variations in symptoms, language, locale, linguistic diversity, and cultural contexts.
Tropical and infectious diseases (TRINDs) exemplify such an out-of-distribution challenge. Prevalent in global low-resource areas, TRINDs impact 1.7 billion people, disproportionately affecting women and children. Effective management is hampered by critical limitations in surveillance, early detection, accurate diagnosis, management, and vaccination strategies. LLMs hold immense potential for early screening and surveillance through symptom, location, and risk factor analysis. Yet, empirical investigation into LLM performance on TRINDs remains limited, with a scarcity of datasets suitable for rigorous evaluation.
To bridge this critical gap, we introduce sophisticated synthetic personas and advanced benchmark methodologies tailored for out-of-distribution disease subgroups. Our TRINDs dataset comprises over 11,000 manually curated and LLM-augmented personas, covering a wide spectrum of tropical and infectious diseases with detailed demographic, contextual, geographical, linguistic, clinical, and consumer attributes. Preliminary findings from this research were presented at the NeurIPS 2024 workshops on Generative AI for Health and Advances in Medical Foundation Models.
play silent looping videopause silent looping videounmute videomute videoVisualizing the comprehensive development process and benchmarking strategy for the TRINDs dataset.
Pioneering Synthetic TRINDs Personas for Advanced LLM Evaluation
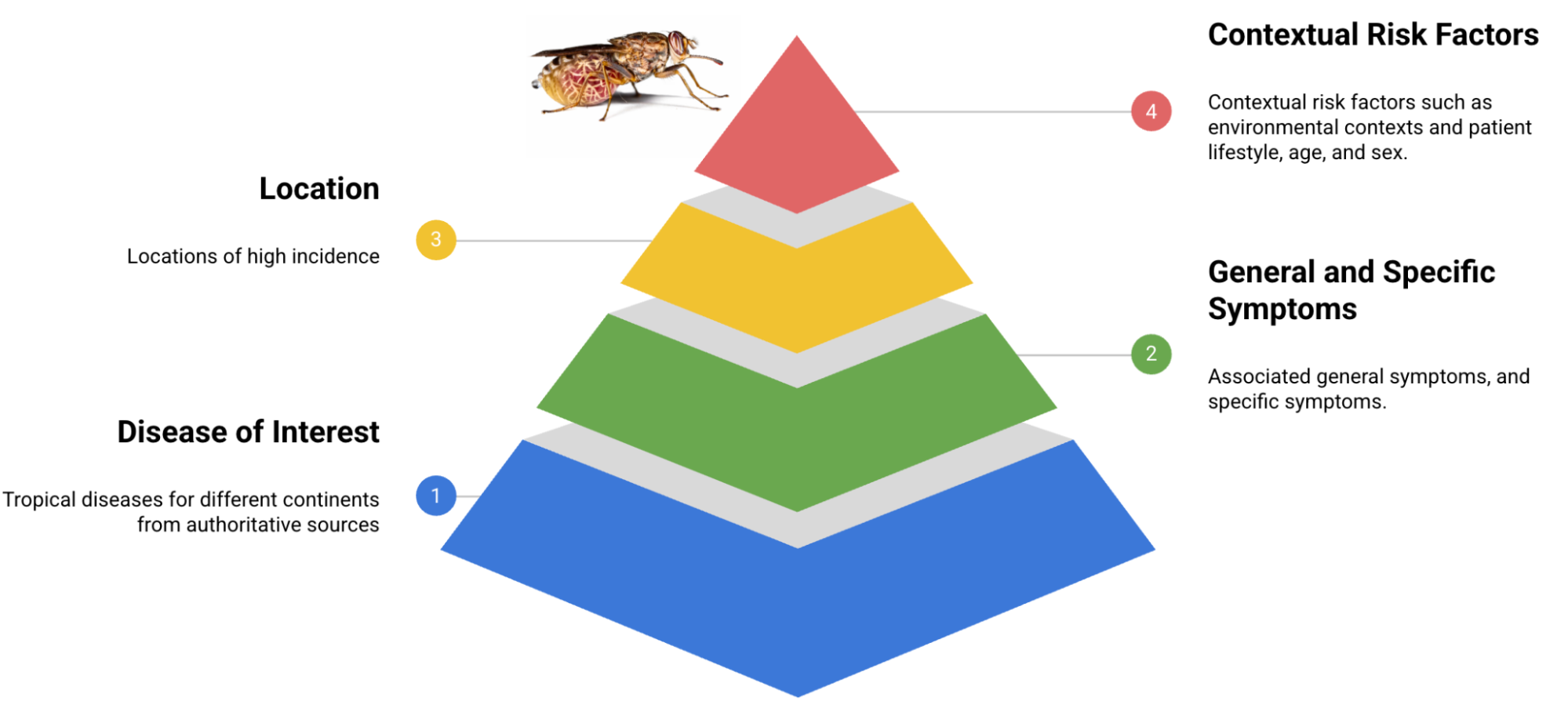
Leveraging authoritative medical resources from global organizations including the WHO, PAHO, and the CDC, we meticulously extracted critical disease information. This knowledge formed the foundation for an initial cohort of patient persona templates, each encompassing general symptoms, direct attributes, and specific manifestations. These personas further integrate contextual, lifestyle, and risk factor elements, rigorously reviewed by clinical experts to ensure accuracy and relevance for persona formatting. This foundational set currently spans 50 distinct diseases.
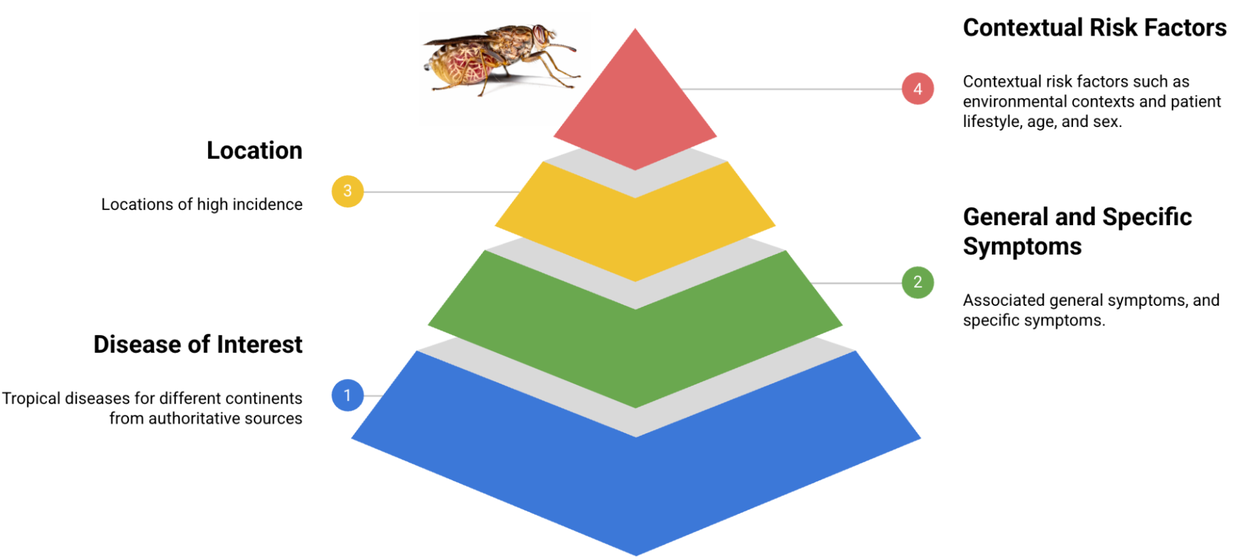
Fundamental components constituting the TRINDs dataset architecture.
Through advanced LLM prompting techniques, we systematically expanded this initial seed set. This expansion incorporated comprehensive demographic and nuanced clinical/consumer augmentations, yielding a dataset exceeding 11,000 personas. To assess language's impact on model performance, we also translated the seed set into French, enabling evaluation across linguistic distribution shifts. Furthermore, we developed an automated LLM-based scoring system that validates an answer's correctness based on the congruence or meaningful similarity between the ground truth and predicted diagnoses.
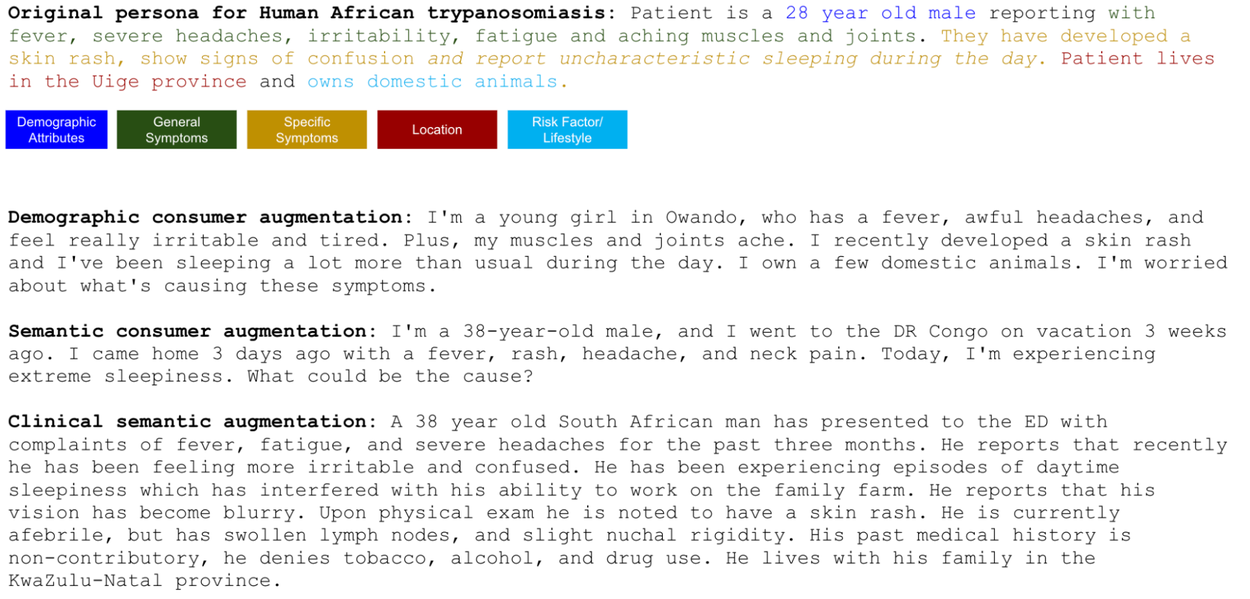
Illustrative examples of an original seed persona alongside LLM-generated augmentations.
Rigorous LLM Performance Evaluation
Comparative Analysis: LLM TRINDs Performance vs. USMLE Benchmarks
We meticulously evaluated the diagnostic accuracy of Google's Gemini models (Gemini 1.5) in identifying diseases based on persona descriptions. Our findings reveal significant distribution shifts, indicating lower performance on the TRINDs dataset compared to reported results on USMLE-based benchmarks.
Contextual Relevance: Impact on Diagnostic Accuracy
We conducted systematic evaluations to discern the influence of diverse contexts, including clinical versus consumer perspectives, demographic variables (age, race, gender), and semantic styles. Our analysis demonstrates that the interplay of symptoms, risk factors, location, and demographics profoundly impacts LLM accuracy in achieving precise diagnoses, underscoring that symptoms alone may be insufficient for reliable diagnostic outcomes.
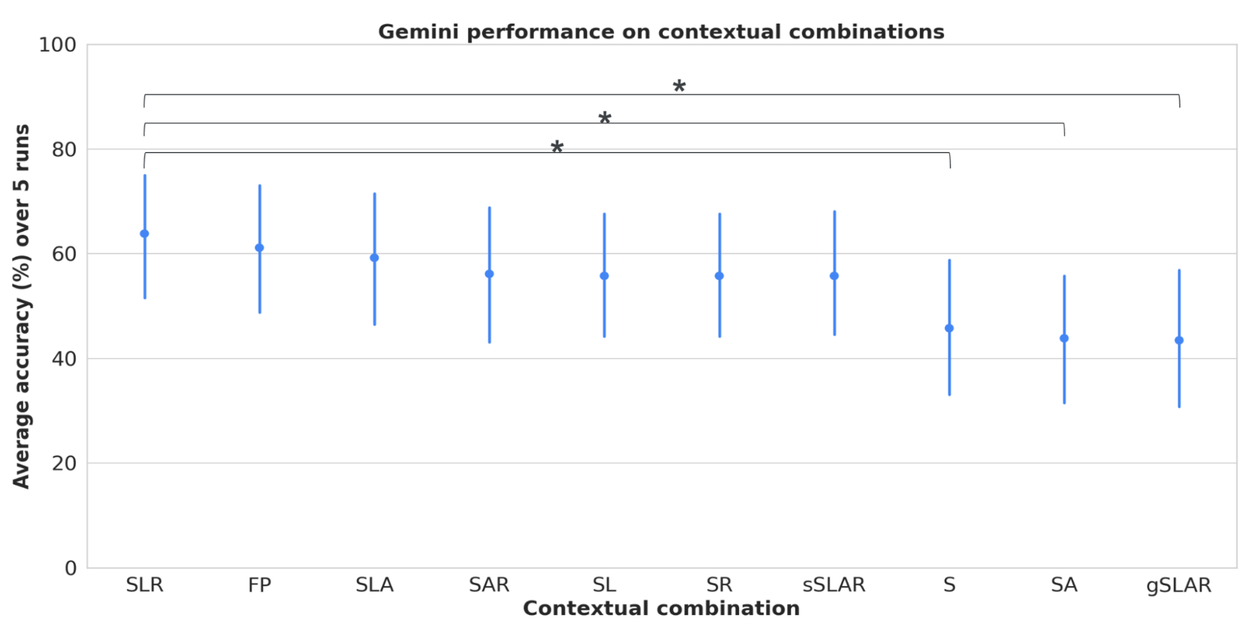
Assessing LLM performance across varied contextual combinations of symptoms (general and specific), geographical location, risk factors, and demographic attributes. S = symptoms, L = location, A = attributes, R = risk factors, gS = general symptoms, sS = specific symptoms, FP = full persona with all context. Error bars indicate the 95% confidence interval. Notably, FP accuracy significantly surpasses S, SA, and gSLAR.
Performance Across Demographic Subgroups and Counterfactual Locations
We rigorously investigated the influence of explicitly stating race (e.g., “I am Black” or “Patient is racially Asian”) and varied gender references (female, male, non-binary) on model performance. Our analysis revealed no statistically significant performance disparities across racial and gender subgroups. We also explored the impact of indicating low-incidence locations (counterfactual locations) on diagnostic accuracy, as detailed in our research paper.
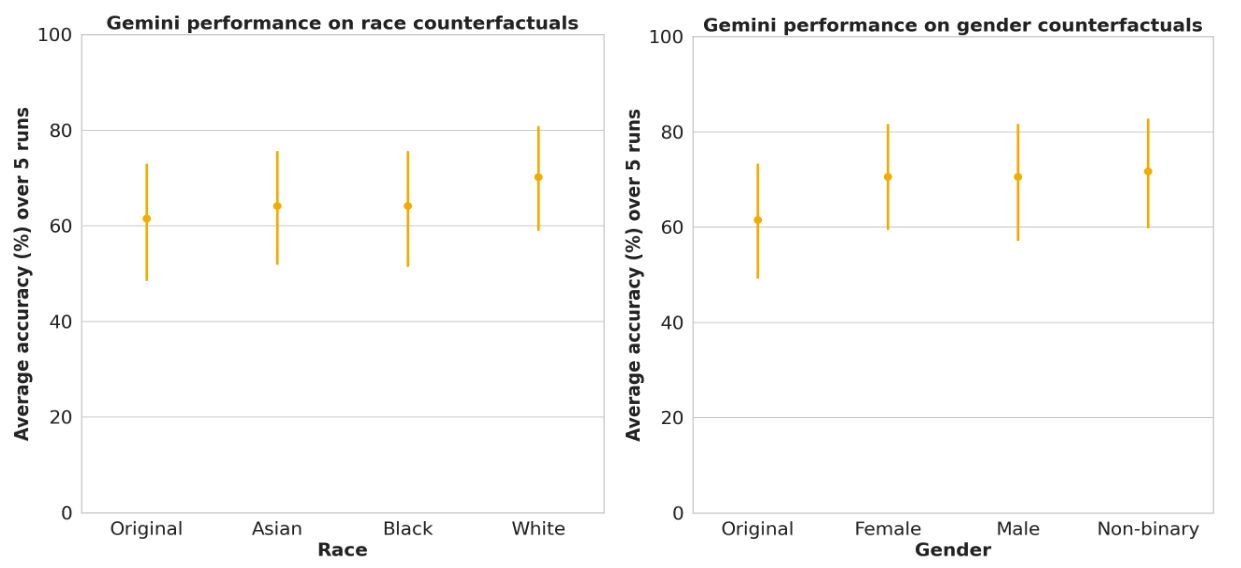
LLM performance across race and gender subgroups within the TRINDs dataset shows no significant performance variance. Error bars represent the 95% confidence interval.
Human Expert Performance Benchmarking and Evaluation
We engaged seven seasoned experts, each possessing over a decade of experience in TRINDs and public health, to address short-answer and multiple-choice questions. We meticulously characterized the performance of the top five experts and simultaneously benchmarked LLM performance on the identical dataset subset. This comparative analysis enabled the simulation of diverse expert consensus scenarios.
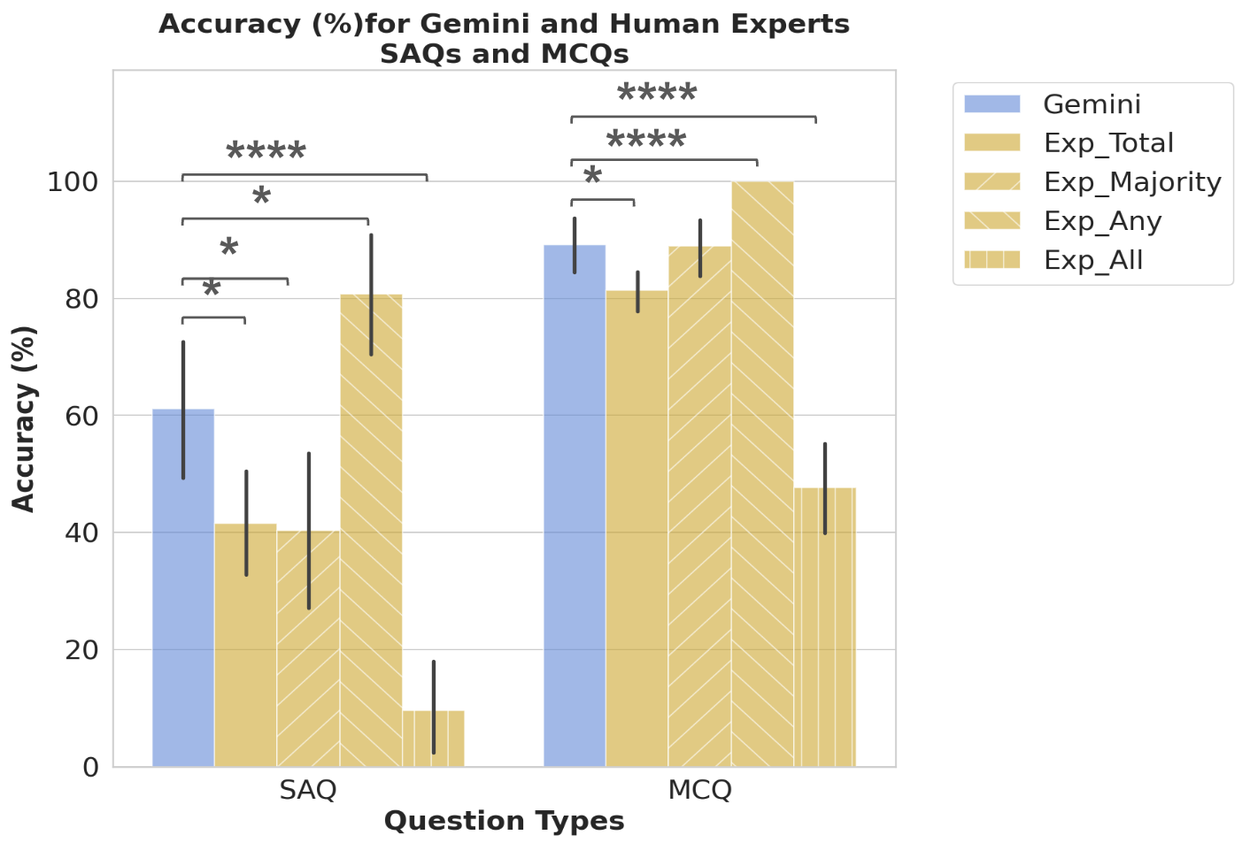
Comparison of LLM (Gemini) performance against top human experts, evaluated using four metrics: 1) average expert score (Exp_Total); 2) full score if the majority vote was correct (Exp_Majority); 3) full score if any expert was correct (Exp_Any); and 4) full score only if all experts were correct (Exp_All). This methodology allows exploration of various expert decision-making dynamics. Error bars represent the 95% confidence interval.
While LLMs exhibited lower performance on TRINDs compared to USMLE benchmarks, they outperformed the highest-scoring individual expert and most expert combination scenarios. The 'Exp_Any' scenario, reflecting a consensus from diverse public health experts, presents a more challenging yet appropriate benchmark that LLMs are beginning to surpass.
Granular Performance Analysis: Per-Disease Accuracy
Our findings indicate that LLMs achieve higher accuracy in identifying common diseases (e.g., HIV) or those characterized by distinct symptoms and risk factors (e.g., rabies). Conversely, certain conditions, such as tapeworm infections, are more prone to misclassification when solely based on symptom presentation. Additionally, disease robustness to counterfactual locations varies, with rabies being less susceptible than some other conditions.
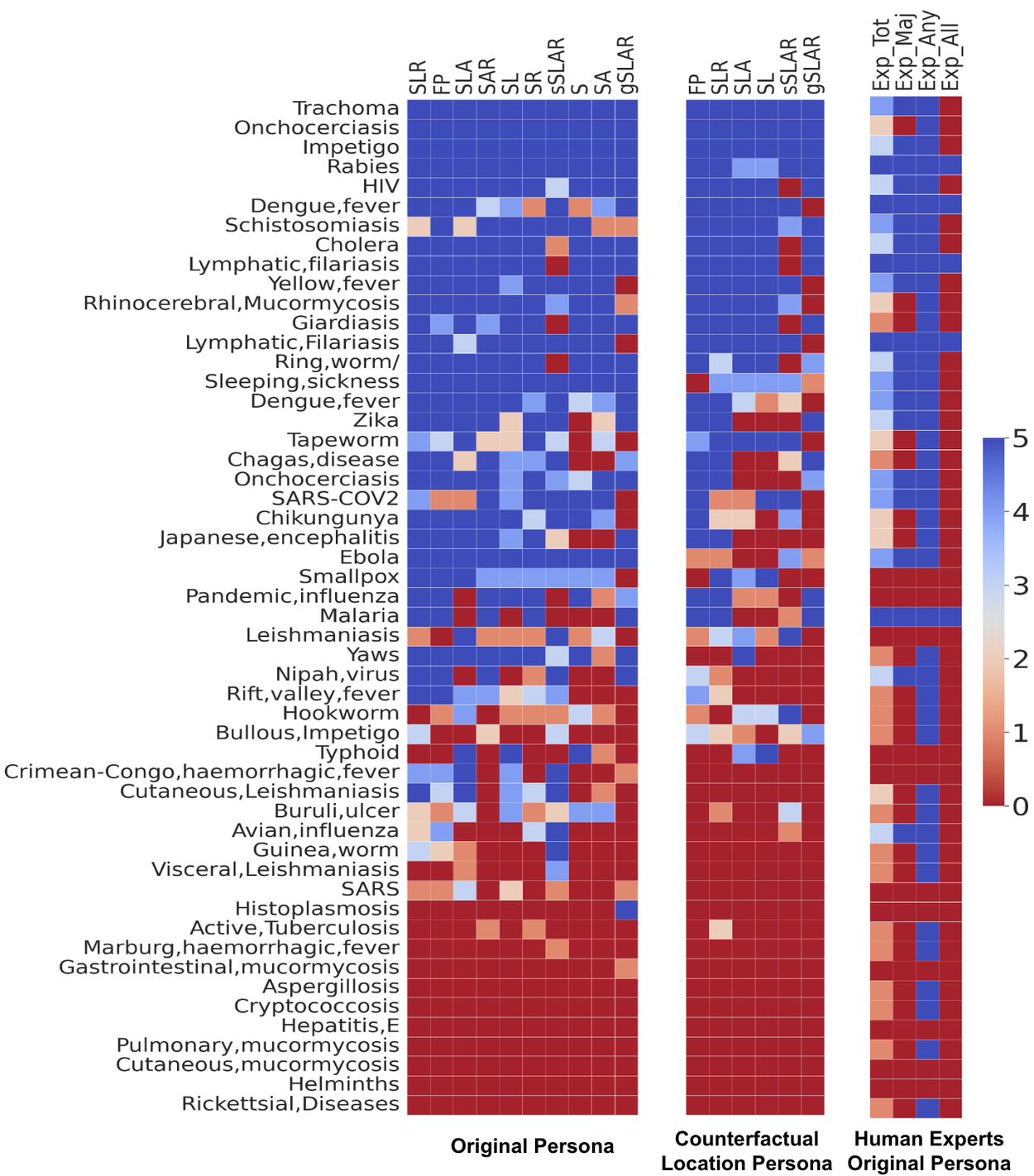
Comprehensive per-disease LLM performance analysis across various contextual combinations (left), including location counterfactuals (center), and comparison with human expert performance (right). Contextual combinations are abbreviated as previously defined.
Enhancing LLM Efficacy via In-Context Learning
We implemented in-context learning by fine-tuning the Gemini 1.5 LLM using multi-shot prompting with the foundational set of 50 seed questions (one per disease), incorporating complete symptom, location, and risk factor data. This strategic tuning significantly improved performance across both demographic and semantic augmentations, demonstrating that existing gaps can be effectively addressed through targeted optimization and that seed datasets are instrumental for enhancing LLM capabilities.
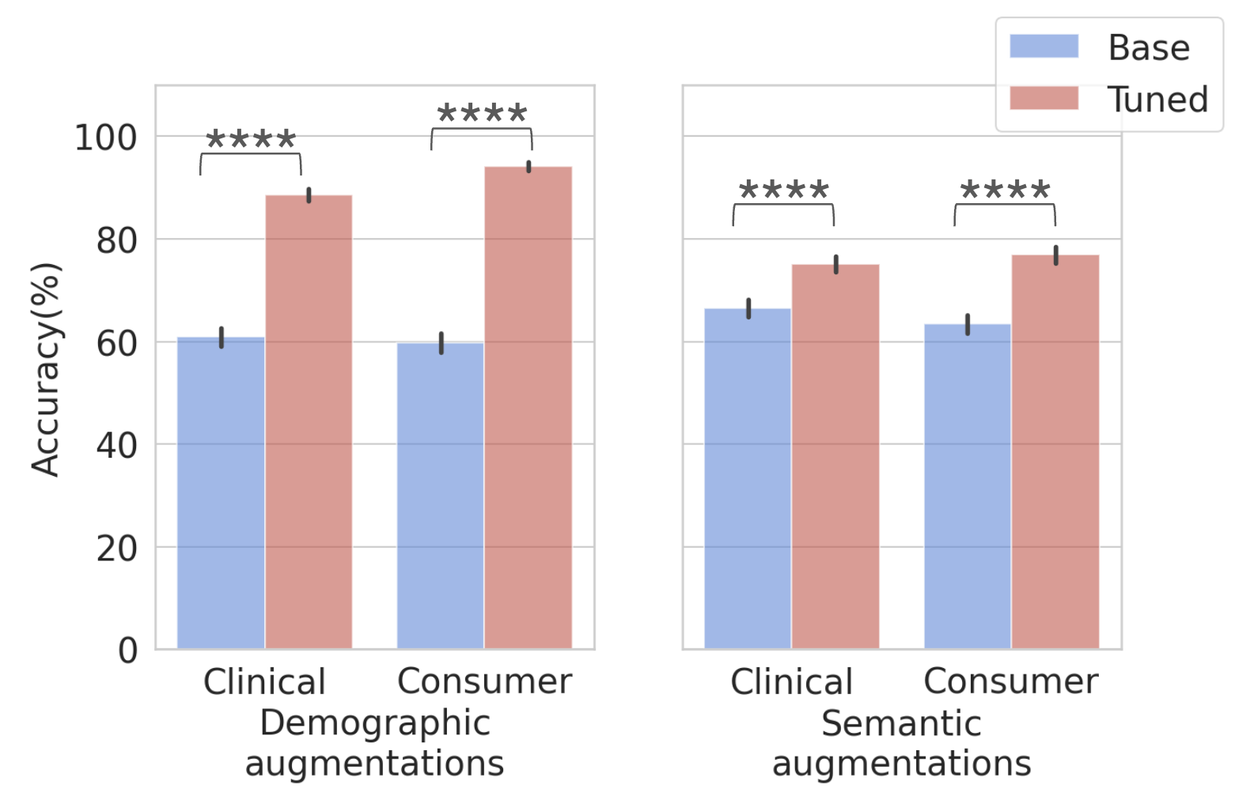
Performance comparison on demographic (left) and semantic (right) augmented datasets pre- and post-in-context tuning. Dataset sizes: Total = 10,570; clinical demographic (n=2635), consumer demographic (n=2635), clinical semantic (n=2650), and consumer semantic (n=2650). Error bars indicate the 95% confidence interval.
Assessing the Potential of LLM-Powered Disease Screening Tools
We have developed an interactive TRINDs disease screening user interface, powered by a specialized Gemini model. This intuitive tool allows users to input demographics, location, lifestyle details, and select symptoms from an extensive list to receive a diagnostic suggestion. Integrated with the Our World in Data API, the interface also displays relevant disease incidence rates.
play silent looping videopause silent looping videounmute videomute videoDemonstrating the user interface and a practical use case for the LLM-powered tool designed for tropical and infectious disease screening.
Preliminary expert ratings indicate that this interface, despite its streamlined output, possesses substantial potential as an impactful and user-friendly reference tool for infectious diseases, offering significant advantages to both clinicians and researchers.
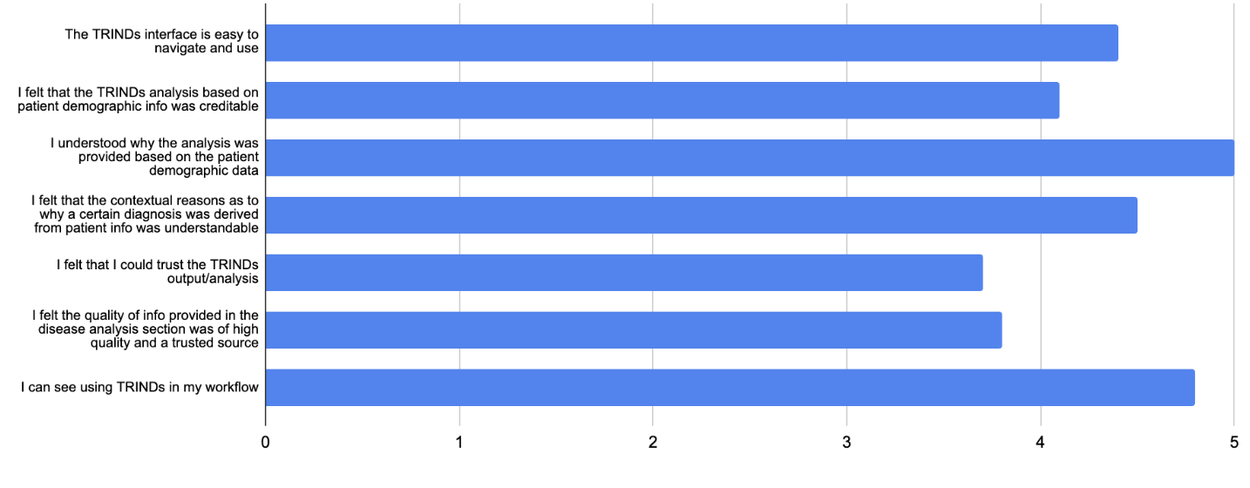
Expert evaluations of the TRINDs disease screening tool.
Key Implications for Global Health
Our research underscores the transformative potential of LLM-driven tools in supporting healthcare workers, particularly in resource-constrained environments. These advanced systems should function as complements to, rather than replacements for, expert clinical judgment. Continuous validation and regular updates are imperative to address evolving real-world scenarios and ensure reliability across diverse clinical settings. Given the critical nature of health outcomes, it is paramount that LLMs are rigorously evaluated for accuracy, contextual understanding, and cultural sensitivity. Transitioning this approach into a clinical tool necessitates further validation and adherence to stringent regulatory review processes. Future research will focus on expanding these benchmarks to incorporate multilingual and multimodal capabilities.
Acknowledgements
We extend our sincere gratitude to the dedicated authors and contributors of this project: Mercy Asiedu, Nenad Tomasev, Tiya Tiyasirichokchai, Chintan Ghate, Awa Dieng, Katherine Heller, Mariana Perroni, Divleen Jeji, and Heather Cole-Lewis. Special thanks to our external expert panel: Oluwatosin Akande, Geoffrey Siwo, Steve Adudans, Sylvanus Aitkins, Odianosen Ehiakhamen, and Eric Ndombi. We also acknowledge the invaluable support and leadership of Marian Croak.


