Cancer is a genetic disease. The controls for cell division go wrong. Many kinds of cancer exist. Each has its own genetic causes. Studying cancer means finding the gene changes in tumor cells. This is key to creating a treatment plan. Doctors now often check the genes of tumor cells. This helps them make plans that stop the cancer from growing.
Our new paper in Nature Biotechnology shows a new tool called DeepSomatic. We worked with partners at the University of California, Santa Cruz Genomics Institute and other researchers. DeepSomatic uses machine learning. It finds gene changes in tumor cells more accurately than older methods. It works with data from all major sequencing tools. It can handle different sample types. It can also learn about new cancer types it was not trained on.
We made the tool and its training data public. This helps the research community. This work is part of Google’s efforts to use AI to understand cancer. This includes AI for screening mammograms for breast cancer. It also includes AI for screening CT scans for lung cancer. We also partner to use AI for research on female reproductive system cancers. Our goal is to speed up cancer research. We want to help make precision medicine a reality.
Gene changes after birth
Genome sequencing finds gene changes. It compares them to the standard human genome. It is hard to tell real changes from errors in sequencing. About ten years ago, Google Research made DeepVariant. This tool finds inherited gene changes. These are called germline variants. They come from parents and are in all body cells.
Cancer genetics are more complex. Cancers often have gene changes that happen after birth. Things like UV light or chemicals can damage DNA. Errors during DNA copying can also cause changes. These changes can happen in somatic cells. Sometimes, these changes make cells grow when they should not. This starts cancer. It can also make cancer grow faster and spread.
Finding changes in some somatic cells is harder than finding inherited changes. Tumor cells can have many different changes. The sequencing error rate can be higher than the rate of a real change in the sample.
Training DeepSomatic to find gene changes in tumor cells
We made DeepSomatic to solve these problems. It finds somatic changes accurately. Most of the time, scientists get tumor cells from a biopsy. They also get normal cells from the body. DeepSomatic learns from both. It finds changes in tumor cells that are not inherited. These changes can show what is making the tumor grow. DeepSomatic can also work with only tumor cells. This is useful for blood cancers like leukemia. It is hard to get only normal cells from a blood sample. DeepSomatic works for many research and medical uses.
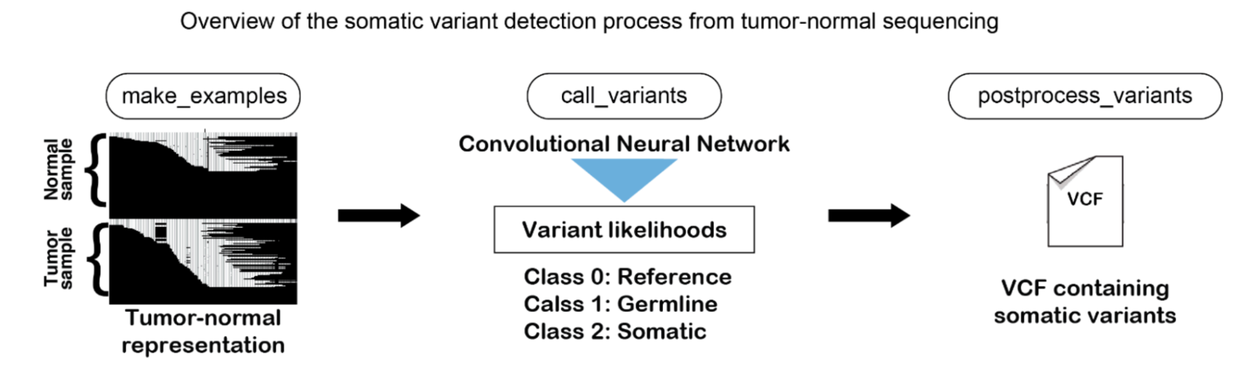
Like DeepVariant, DeepSomatic turns gene sequencing data into images. These images show the sequence data and other details. DeepSomatic uses its convolutional neural network. It looks at data from tumor cells and normal cells. It tells apart the standard genome, the inherited changes, and the cancer changes. It ignores errors from sequencing. The result is a list of cancer-related changes, or mutations.

DeepSomatic finds cancer changes in gene data. First, it turns sequencing data from tumor and normal cells into an image. DeepSomatic uses its convolutional neural network on these images. It tells apart the standard genome, normal gene changes, and cancer changes. It ignores small sequencing errors. This gives a list of cancer-causing changes, or mutations.
To train good models, we need a lot of high-quality data. We created a new data set for finding changes in tumor cells. We worked with UC Santa Cruz and the National Cancer Institute. We sequenced tumor cells and normal cells from four breast cancer samples and two lung cancer samples.
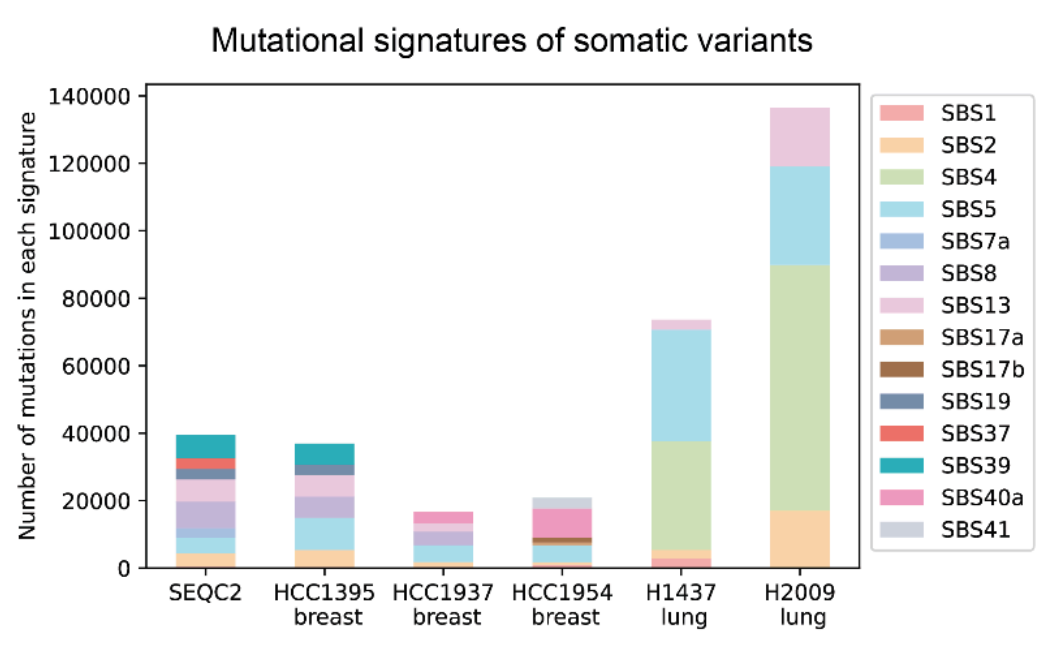
This shows the data used to train DeepSomatic. Each bar shows the number of mutations in six cancer samples. Color shows different mutation types. Lung cancer shows a type of mutation caused by toxins. But even the same cancer type has big differences. These differences can show how well it will react to treatment.
We sequenced these six samples using three main methods: Illumina’s short-read sequencing, PacBio’s long-read sequencing, and Oxford Nanopore Technology’s long-read sequencing. We combined data from all three. This removed errors unique to each platform. This created an accurate data set called the Cancer Standards Long-read Evaluation dataset (CASTLE).
Testing DeepSomatic’s ability to find cancer changes
We trained DeepSomatic on three breast cancer genomes and two lung cancer genomes from the CASTLE data. We then tested DeepSomatic on one breast cancer genome not used in training. We also tested it on a part of each sample’s chromosome 1, which was also left out of training.
Results show DeepSomatic models worked better than other methods. They found more tumor variants with higher accuracy. We compared DeepSomatic to SomaticSniper, MuTect2, and Strelka2 for short-read data. For long-read data, we compared it to ClairS. DeepSomatic uses a deep learning model trained on real data.
DeepSomatic found 329,011 somatic variants in the six samples. It is especially good at finding cancer changes with insertions and deletions (Indels). For these, DeepSomatic greatly improved the F1-score. The F1-score measures how well a model finds true variants without finding fake ones. On Illumina data, the next best tool scored 80% for Indels. DeepSomatic scored 90%. On Pacific Biosciences data, the next best scored under 50%. DeepSomatic scored over 80%.

DeepSomatic results (purple) for a breast cancer sample compared to other tools. Some tools work for Illumina data. Only one other tool (pink) works for long-read data from PacBio and Oxford Nanopore. The F1-score shows how many variants are found and how accurate they are. DeepSomatic is slightly better for single-letter changes (single nucleotide variations). It is much better for Indels.
The seventh sample was a breast cancer tumor preserved with formalin-fixed-paraffin-embedded (FFPE). This common method can damage DNA. This makes genetic analysis harder. This sample was also sequenced using whole exome sequencing (WES). WES focuses on the 1% of the genome that makes proteins. DeepSomatic trained on this type of data. It tested on chromosome 1. It again performed better than other tools. This shows it can find variants in older or lower-quality samples. It can also work on data where only the exome was sequenced.

DeepSomatic is more accurate on samples with difficult pre-processing. This includes fixed formalin paraffin embedded (FFPE), a way to keep tissue samples (left). It also includes whole exome sequencing (WES), which sequences only the protein-coding parts of the genome (right). The middle section shows a sample with both FFPE and WES.
Applying DeepSomatic to other cancers
We tested DeepSomatic on other cancers. We looked at a glioblastoma sample. This is a fast-growing brain cancer. It starts with a few gene changes. DeepSomatic found those changes. This shows it can learn and work for other cancer types.
We also worked with Children’s Mercy to study pediatric leukemia. This is a common cancer in children. Leukemia is in the blood. So, a normal blood sample is not possible. DeepSomatic found the known variants. It also found 10 new ones. This shows it can work with only tumor samples.
What’s next
We hope labs and doctors can use this tool. Finding known cancer variants could help choose treatments. This includes chemotherapy or immunotherapy. Finding new variants could lead to new therapies. We hope people can use these tools to learn more about tumors. They can find what drives the cancer. Then they can give patients the best treatments.


