
Harnessing unprecedented data streams from advanced medical technologies to personal smartwatches empowers researchers and clinicians. Electronic health records, medical imaging, diagnostic tests, genomic data, and wearable sensor measurements generate rich datasets. These diverse data streams contain unique and overlapping signals, even within the same organ system. For instance, in the cardiovascular system, electrocardiograms (ECG) measure electrical activity, while photoplethysmograms (PPG), common in smartwatches, track blood volume changes. Simultaneously analyzing these modalities offers a comprehensive view of heart health, assessing both electrical function and pumping efficiency. Integrating these physiological signatures with genomic data from large-scale biobanks accelerates the identification of disease-related genetic factors. Our prior research, REGLE, successfully performed genetic discovery using single data types. However, analyzing each modality separately (Unimodal REGLE or U-REGLE) may miss subtle shared information. We hypothesized that jointly modeling complementary data streams amplifies biological signals, reduces noise, and drives more powerful genetic discoveries. This paper presents our latest findings in the *American Journal of Human Genetics*: "Utilizing multimodal AI to improve genetic analyses of cardiovascular traits." We introduce M-REGLE, a multimodal extension of REGLE that analyzes multiple clinical data types concurrently. M-REGLE achieves lower reconstruction error, identifies more genetic associations, and significantly improves cardiac disease prediction compared to U-REGLE.
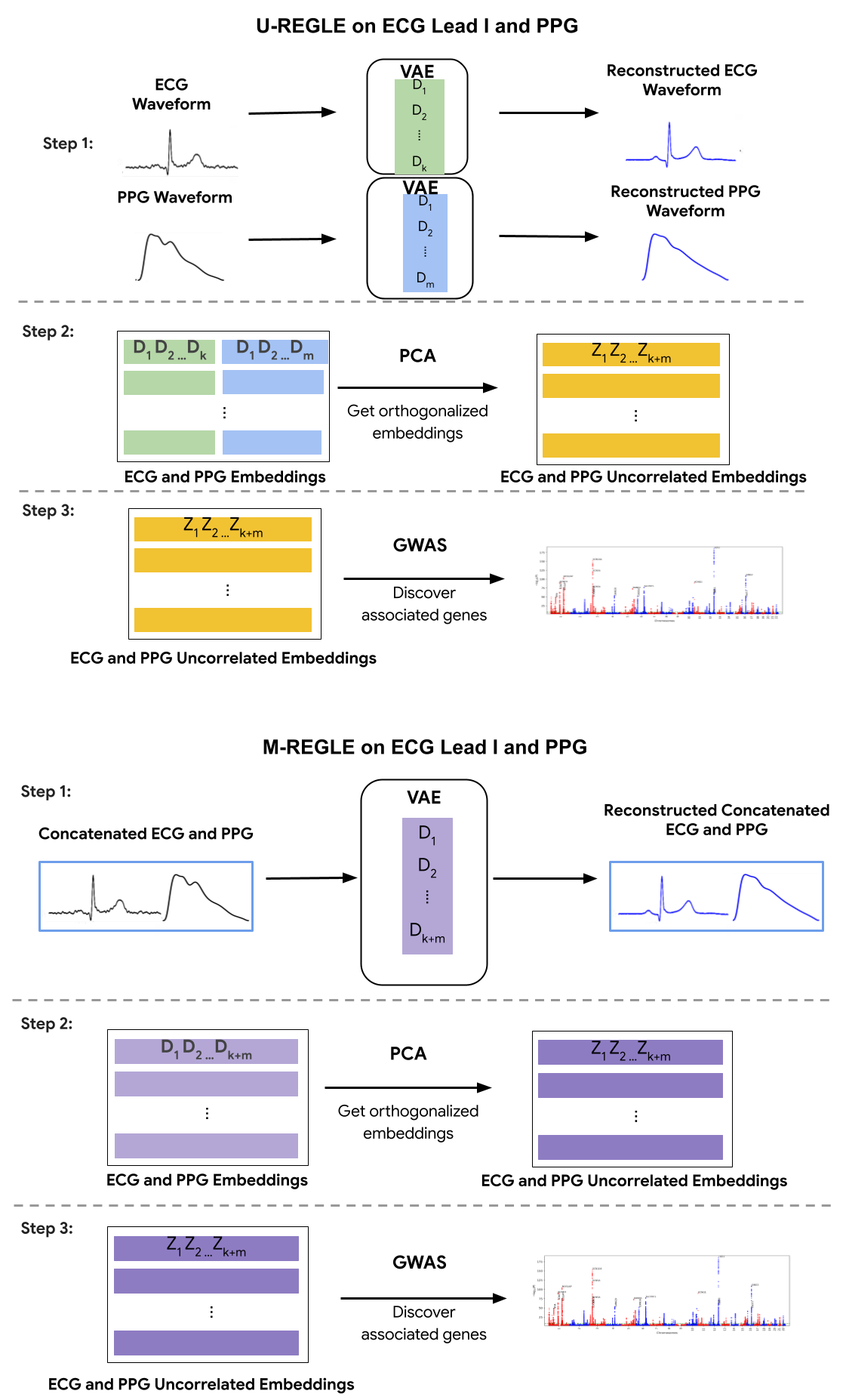
M-REGLE overview demonstrates its steps compared to running our previous model REGLE on each modality separately (U-REGLE, or Unimodal REGLE).
The challenge: Seeing the whole picture
M-REGLE's core principle is that diverse clinical modalities, particularly within the same organ system like the circulatory system, encode both complementary and overlapping information. For example, the 12 leads of an ECG provide distinct body surface perspectives. Physicians analyze specific leads to pinpoint heart attack locations or diagnose arrhythmias. M-REGLE's integrated approach, combining multiple modalities like the 12 ECG leads or ECG with PPG data before representation learning, yields a more accurate tool that excels at identifying genetic associations, analyzing complex physiological data, and predicting disease. M-REGLE employs a robust, multi-step joint learning process. Instead of analyzing individual ECG leads or ECG and PPG waveforms separately, M-REGLE first merges them. It then utilizes a convolutional variational autoencoder (CVAE) to learn a compressed, combined "signature"—latent factors—from these multiple data streams. The CVAE compresses essential information into a lower-dimensional, largely uncorrelated representation through encoder and decoder networks. The encoder maps ECG and PPG waveforms to latent factors, while the decoder reconstructs waveforms from these factors. Principal component analysis (PCA) is applied to the CVAE-generated signatures to ensure learned factors are truly independent. Finally, we identify associations (significant correlation) between these computed independent factors and genetic data using genome-wide association studies (GWAS). Individual GWAS results are statistically combined to pinpoint genetic variations linked to the underlying physiological system.
Better learned representations
M-REGLE significantly advances U-REGLE by consistently generating superior "learned representations" of the data. Medical data, such as an ECG, comprises hundreds of individual data points. When analyzing multiple medical modalities, M-REGLE condenses the most critical characteristics into "latent factors" rather than processing each modality independently. This approach dramatically reduces reconstruction errors and more effectively captures essential information from the original waveforms compared to unimodal learning. For 12-lead ECGs, M-REGLE achieved a 72.5% reduction in reconstruction error.
Interpretability sheds some light on embeddings
Generative AI's interpretability is a key advantage. Our study leverages M-REGLE embeddings to demonstrate the relationship between these embeddings and ECG and PPG waveforms. Specifically, we show how altering individual embedding coordinates affects the reconstructed ECG and PPG waveforms via the M-REGLE decoder. We focused on identifying coordinates that effectively distinguish between samples with and without atrial fibrillation (AFib). M-REGLE embeddings at positions 4, 6, and 10 proved most distinctive. Modifying the 4th M-REGLE embedding values within [-2, 2], while keeping other embeddings constant, produced observable changes in the reconstructed ECG lead I and PPG. The T-wave segment of ECG lead I changed in magnitude, and the PPG signal's dicrotic notch showed a minor alteration. The dicrotic notch provides critical information about cardiovascular function and health; a less prominent or absent notch often indicates increased arterial stiffness.
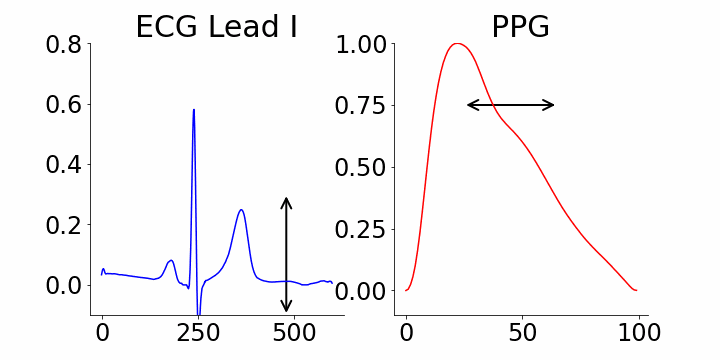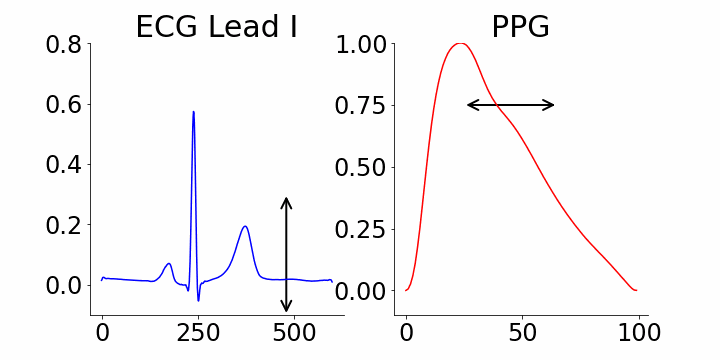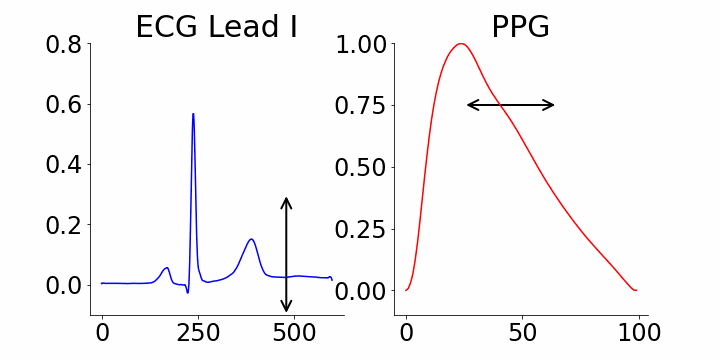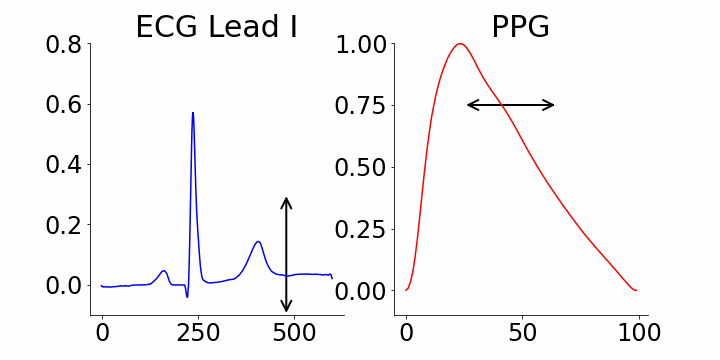
Varying the 4th M-REGLE embedding impacts the reconstructed ECG Lead I and PPG, reducing the magnitude of the T-wave segment in ECG lead I (left) and altering the prominence of the dichroic notch in the PPG (right).
Enhanced genetic discovery
M-REGLE significantly improves upon U-REGLE in identifying genetic associations with cardiovascular disease. For 12-lead ECGs, M-REGLE discovered 19.3% more associated genetic loci (genomic regions) than the unimodal approach. Analyzing ECG lead I plus PPG data, M-REGLE identified 13.0% more loci. Crucially, a substantial majority of these discoveries (24/35 for 12-lead ECG and 11/12 for ECG lead I + PPG) replicated known genetic associations for ECG or PPG traits as documented in the GWAS catalog. M-REGLE also revealed several novel loci not previously linked to these traits, some of which demonstrated connections to cardiovascular traits in other databases.

This 3-way Venn diagram compares GWAS catalog loci, loci discovered by M-REGLE (12-lead ECG), and loci discovered by U-REGLE. The GWAS catalog represents previously identified loci.
Improved polygenic risk scores
A polygenic risk score (PRS) quantifies an individual's genetic predisposition to a disease. We found that PRS developed using genetic variants identified by M-REGLE (from 12-lead ECG data) significantly outperformed those derived from U-REGLE in predicting cardiac disease, particularly atrial fibrillation (AFib). M-REGLE's PRS demonstrated superior accuracy in identifying individuals at risk. These improvements for AFib were consistently observed not only in the UK Biobank but also independently validated across other large datasets, including the Indiana Biobank, EPIC-Norfolk, and the British Women's Heart and Health Study.
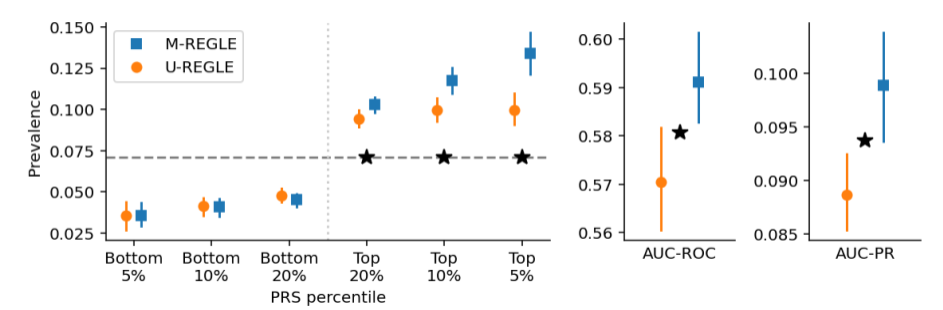
This comparison of M-REGLE PRSs for atrial fibrillation (AFib) shows prevalence, AUC-ROC, and AUC-PR computed (* indicates a statistically significant difference).
Why does M-REGLE work?
M-REGLE's effectiveness stems from its sophisticated information handling. By simultaneously considering multiple modalities, M-REGLE achieves three primary benefits. First, it efficiently captures shared information, learning it once across all relevant modalities. Second, it amplifies unique and complementary signals inherent to each modality. Third, M-REGLE effectively reduces noise, as information from one modality can clarify or filter noise in another. Collectively, these advantages produce a clearer, more robust signal, enabling powerful downstream genetic analysis.
The future is multimodal
This research marks a significant advancement in utilizing the increasingly abundant, multimodal health data. M-REGLE provides a method to uncover novel genetic links to complex diseases, enhance disease risk prediction capabilities, and potentially identify new therapeutic targets. Furthermore, with the proliferation of smart wearables continuously collecting physiological data like ECG and PPG, methods such as M-REGLE are essential for translating health data into actionable insights and, ultimately, achieving better health outcomes.
Acknowledgements
This project represents a collective effort from numerous contributors and institutions. We express our sincere gratitude to our collaborators for their invaluable contributions: Yuchen Zhou, Justin Cosentino, Howard Yang, Andrew Carroll, Cory Y. McLean, Babak Behsaz (Google); Zachary R. McCaw (University of North Carolina); Tae-Hwi Schwantes-An, Dongbing Lai (Indiana University); Mahantesh I. Biradar, Robert Luben, Jorgen Engmann, Rui Providencia, Anthony P. Khawaja (University College London); Patricia B Munroe (Queen Mary University of London). We also thank Anastasiya Belyaeva for manuscript review, Greg Corrado, Shravya Shetty, and Michael Brenner for their support, and Monique Brouillette for her assistance in crafting this blog post.


